Supplementary material for “Human error in high-biocontainment labs: a likely pandemic threat”
By Lynn Klotz | February 25, 2019
Categorizing human errors. Because of the scarcity of data on human-error incidents in high-biocontainment labs, Gryphon Scientific looked to other sectors to define and understand types of human error[i]. Gryphon summarized its findings in a table[ii] in its report. Gryphon lists three types of human error, summarized here in Table 1 in shortened and modified form.

Table 1. Types of human error. According to Gryphon, much of the data on human reliability comes from the transportation, chemical and nuclear sectors. PPE stands for personal protective equipment.
__________________________________________________________________________
Many different data sources confirm the idea that human error is the main source of incidents in BSL3 and BSL4 laboratories. Here the data from various sources are summarized, with detailed analysis available from the author.
FSAP/CDC Incident Reports. The following incident data are from the Federal Select Agent Program (FSAP) yearly summary reports to Congress[iii] for seven years, 2009 through 2015. During that period there were a total of 749 incidents reported to FSAP from select-agent research laboratories. In December 2016, the percent of BSL3 registered select-agent labs was about 70 percent[iv] of all labs, a ratio here assumed to be relatively constant over the seven-year period.
The data on incidents were provided to Congress in seven categories listed in Table 2.
Table 2. Numbers of incidents for the seven categories reported to Congress by FSAP/CDC for the years 2009 through 2015. The five categories due to human error are highlighted in bold-face type. BSC stands for biosafety cabinet.
_____________________________________________________________________
The 79.3 percent human error supports the premise of this analysis.
The bold-faced categories are likely close to 100 percent human error. Category 4 could be a mix of human error and personal protective equipment (PPE) defects or failure. To be conservative, assume Category 4 is entirely PPE defects or failure. The largest incident category is spills, Category 6, a category where some causes could be reduced (see below).
By FSAP rules[v], spills or splashes or other accidents in a BSL3 laboratory where the worker wears personal protective equipment (PPE) including a powered air purifying (PAPR) or the incident occurred in a biosafety cabinet (BSC) are not considered reportable incidents since the worker is not exposed to pathogen. On the other hand, all spills, splashes, etc. are reportable to NIH[vi] regardless of whether the worker is protected by PPE or a BSC. The FSAP rules seem more reasonable.
NIH incident reports. The FOIA-requested incident reports from the NIH Office of Science Policy cover the period from 2004 through 2017[vii] and cover BSL3 and BSL4 labs. There were no reported incidents from BSL4 labs. Reporting to NIH is required only for incidents involving pathogens that contain recombinant DNA. While there have been incidents in BSL4 labs, they may not have involved strains containing recDNA so would not show up in the FOIA NIH reports. BSL2 labs must report incidents as well, but they are of less interest here; thus, the very many BSL2 reports were not requested.
Recombinant DNA research is ubiquitous in molecular biology; and sometimes, pathogens are engineered to contain recDNA. Thus, a wider group of pathogens than FSAP Select Agents would be covered in the FOIA NIH reports. However, there is overlap in reports to FSAP and NIH.
The BSL3 reports provide extremely detailed descriptions of incidents from the facilities where the incidents occurred. The reports are often several dozen pages long, so almost no questions remain about details, and much can be learned about incidents and human error.
The 123 FOIA NIH reports cover 128 incidents. A few reports cover more than one incident. In the 128 incidents, 86 are due to human error, yielding 67.2% percent human error. The master spreadsheet with facility names removed is pictured in Appendix 1, since some facilities have claimed confidentiality for their reports. While the reports are actually public records, their identity is not necessary for documentation here. Appendix 1 can be employed to calculate all sorts of useful numbers, such as percentage of incidents that result in undetected or unreported laboratory-acquired infections (uuLAIs). Since workers with uuLAIs will leave the facility after work, these infections represent a release into the community, so they are key data for analyzing risk of a community outbreak.
Since reporting to NIH in past years has been sporadic, it is likely that many incidents had not been reported. Even in recent years, there is no guarantee that most incidents have been reported despite NIH’s efforts to have all recombinant DNA labs report. In general, the analysis here is accomplished using methods that don’t require identifying facilities that haven’t reported incidents. In contrast, since the FSAP has strict requirements for compliance to Select Agent rules and enforced by the FBI, the reporting percentage is likely high.
In the particular case of human error, low reporting rate is unlikely to affect the percentage, since % human error = (number of incidents due to human error)/(number of incidents). Both the numerator and denominator are likely changed by the same factor by non-reporting.
From the analysis of the spreadsheet pictured in Appendix 1, most human error is skill-based:
- skills error = 68.2 percent
- rules error = 25.0 percent
- knowledge error = 3.4 percent
- % other error = 3.4 percent
This is consistent with the FSAP data.
The percentage of types of skill-based errors from the FOIA NIH data are:
- Potential needle stick or other potential through the skin exposures with other contaminated objects = 47.1 percent
- Spills, splashes, dropped containers = 42.6 percent
- Bite/scratch from infected animal = 7.4 percent
- Centrifuge, shaker, etc. that is, releases that could have generated an aerosol =2.9 percent
By far, lab workers with potential through the skin exposures and spills-splashes or dropped-objects are the main causes of human-error incidents. For the few incidents relating to centrifugations and shakers, most are caused by defective labware such as cracked centrifuge tubes, cracked shaker flasks, or other defective labware, rather than equipment failure.
Some human errors are “one-off” errors, meaning they happened once, likely won’t happen again, and would be difficult to anticipate. It is unlikely that one can devise meaningful changes in standard operating procedures (SOPs) for many of them for future prevention. Here are a few examples of potentially one-off errors gleaned from the first few-dozen reports:
1) A spill of animal bedding potentially contaminated with recombinant SARS Coronavirus. Cages were stacked next to a -80° C freezer, and when the door was opened a cage tipped over and fell on the floor. This caused the lid of the cage to open and contaminated bedding to fall onto the floor.
2) A water overflow from an effluent decontamination system (EDS) which handles all the effluent from three high-containment suites in a multi-lab BSL3 facility. Water from the EDS backed up in the BSL- 3Ag suite and, to a lesser extent, the ABSL-3 suite. The backup was primarily caused by a staff member’s failure to turn off the sink faucet in the BSL-3 suite, and the EDS could not handle the volume of water produced. A secondary cause was the failure of a high-water-level alarm linked to the EDS to be sent to the appropriate maintenance staff. The water came out of floor drains within the high-containment suites. The BSL-3 suite is located on the main floor above the BSL-3Ag and ABSL-3. While the water produced in the BSL-3 was the cause, this suite was unaffected due to its location.
Work conducted in one room involves 1918 H1N1 viruses.
3) A graduate student sustained a superficial abrasion to the right forearm when his hand slipped while closing the sash of a cage change station.
4) A researcher was exchanging two plastic 24-well plates in the tabletop Sorvall centrifuge. While closing the lid, it was caught on a centrifuge wrench which was accidently placed into the path of the lid. The wrench jumped and knocked one of the removed 24-well plates onto the counter. The plate landed at approximately a 45-degree angle and lost approximately half its contents to the bench top.
5) A researcher was working in a BSC and was transferring inactivated HIV into an ultra-centrifuge tube to layer it over a sucrose gradient. The tube was in a small beaker, and was at an angle to facilitate adding the virus suspension. The tube moved slightly; and to prevent the tube from spilling, he used his left hand to steady it. During this motion, he accidentally moved his right hand, which was holding the syringe, and brushed the tip of the needle over the surface of his left thumb. He felt a small bump or prick, as the needle punctured both gloves on the thumb of the left hand.
6) A researcher was using an electroporator to transform M. tuberculosis cells when the cap of the electroporation cuvette ejected from the machine and landed on the floor, resulting in a minor spill.
Other errors are frequent, for which there are now changes in SOPs in some facilities to reduce their frequency. For instance, needle sticks can occur using syringes with sharp-metal needles to transfer liquids from one small container to another. For injecting animals, sharp-metal needles are needed; but for liquid transfers, blunt-plastic needles would suffice.
Another example: dropped items sometimes could be prevented using lab carts to transport items from place to place instead of carrying them by hand. Some facilities have changed their SOPs to reflect these ideas.
Incident category definitions in the Gryphon influenza lab data are not straight forward for quantifying human error, so percent human error for these influenza labs cannot be determined with confidence.
Failure to inactivate. Failure to inactivate is a major reason for release from high biocontainment into BSL2 labs. Since there are reliable inactivation procedures, failure to inactivate is a human error of the Rules or Knowledge type.
Carrying out research in BSL3 and BSL4 laboratories is difficult, both because of restricted movement in the PPE that must be worn and because of constraints in SOPs to minimize potential exposure to pathogens. It is so much easier to carry out research at BSL2, since researchers are much less constrained. When research does not require active pathogens, researchers will inactivate them so they may be researched at BSL2. If the pathogen has not been inactivated, the threat of a uuLAI increases and the probability of other release into the community increases. In BSL2, workers clothing and body parts are exposed to pathogens.
How often is failure to inactivate responsible for transferring BSL3 and BSL4 pathogens to lower biocontainment? The GAO has weighed in on this question.[viii]
“The total number of incidents involving incomplete inactivation…that occurred from 2003 through 2015 is unknown for several reasons. One key reason is that the [FSAP]…does not require laboratories to identify such incidents on reporting forms. According to the [FSAP…, 10 incidents occurred from 2003 through 2015. However, GAO identified an additional 11 incidents that the program did not initially identify.”
Information about the 11 additional incidents is summarized in the GAO’s Table 3.
The GAO further calls attention to a well-publicized, large incident.
“In May 2015, the Department of Defense (DOD) discovered that one of its laboratories inadvertently sent live Bacillus anthracis, the bacterium that causes anthrax, to almost 200 laboratories worldwide over the course of 12 years. The laboratory believed that the samples had been inactivated…In this case, DOD was inactivating samples to support research on the detection, identification, and characterization of biological threats.”[ix]
The GAO describes yet another well-publicized incident.
“Similar incidents have occurred in other countries, including China, where two researchers conducting virus research were exposed to severe acute respiratory syndrome (SARS) coronavirus samples that were incompletely inactivated. The researchers subsequently transmitted SARS to others, leading to several infections and one death in 2004.”[x]
Thus, failure to inactivate is likely a major path for deadly pathogens to be released from BSL3 and BSL4 containment increasing the probability of release into the community.
BSL4 laboratory releases due to human error. In BSL4 laboratories, researchers don suits that fully cover their bodies, and they breathe outside-air from hoses tethered to them. The mental image is that of astronauts undergoing a spacewalk fully protected from the cold and vacuum of outer space. Thus, researchers would seem to be protected from uuLAIs.
From 1988 until the recent past, there were two direct releases into the community of pathogens (foot and mouth disease virus, Marburg virus) from BSL4 containment, one due to human error. More recently, there have been four releases of Ebola and Marburg viruses from BSL4 to lower containment labs, all due to human error.
While two releases seem small, the number of BSL4 labs is small compared to the number of BSL3 labs. For instance, the FSAP reports about 200 registered BSL3 labs for each of the years between 2009 through 2015, with 6 reported uuLAIs. In comparison, there are now 22 BSL4 labs worldwide[xi]. While the data set is small, the rate of releases from BSL4 labs is comparable to that of BSL3 labs in the U.S.
Here are more details on the two direct releases into the community from BSL4 containment:
(1) In 1990, a 35-year-old junior scientist from the Vektor Laboratory of the Novosibirsk Scientific Center contracted a Marburg virus infection. “[I]n violation of safety regulations, he worked with blood serum of laboratory animals infected with that virus, considering the material to have lost its infectivity in view of its storage at a temperature of 4oC for about 6 months…[F]eeling unwell on April 13th, he went home from work without alerting the laboratory’s medical service, and on April 14th he entertained 10 guests at home. In fact, up until the time he was hospitalized, the patient was in close contact with 12 relatives (his wife and daughter – every day of illness up until hospitalization, and 10 people as guests…)”[xii] He ultimately survived, and it is very lucky no one else was infected.
(2) In August of 2007, “An outbreak of foot and mouth disease was confirmed at a farm in Surrey, U.K…It was concluded that the Foot and Mouth Disease Virus likely originated from the nearby Pirbright Research and manufacturing site in Surrey because of construction activities surrounding a leaking drainage pipe.” [xiii] While the NBAF report says “likely,” the FMDV outbreak almost certainly came from the Pirbright BSL4 laboratory. This is an example of release from a BSL4 facility through an infrastructure design failure, not human error.
On the web site of Boston University’s National Emerging Infectious Disease Laboratories (NEIDL), they argue that the state-of-the-art design and construction of the NEIDL would prevent pathogens from escaping into the community.
“The NEIDL is a 192,000-square-foot, seven-story building designed in accordance with the most stringent and protective measures defined by the National Institutes of Health. It was built on the experience of six existing BSL-4 facilities in North America, none of which has ever had a release or community incident… All critical building systems within the NEIDL have a redundant system to ensure safety and uninterrupted operation at all containment levels.”[xiv]
While there is no problem believing that modern BSL4 labs have been designed and built to the state-of-the-art. While technically correct, there is one misleading phrase in the above quote, namely that there has never been a release from a BSL4 lab in North America. As noted above, the GAO has uncovered four recent releases from BSL4 labs, two of the deadly Ebola virus and one of the deadly Marburg virus.[xv] These three were due to failure to inactivate the viruses before transferring them to a lower BSL2 containment laboratory.
The fourth release in 2014 from the CDC labs occurred when, in the GAO’s words,
“Scientists inadvertently switched samples designated for live Ebola virus studies with samples intended for studies with inactivated material. As a result, the samples with viable Ebola virus, instead of the samples with inactivated Ebola virus, were transferred out of a BSL-4 laboratory to a laboratory with a lower safety level for additional analysis. While no one contracted Ebola virus in this instance, the consequences could have been dire for the personnel involved as there are currently no approved treatments or vaccines for this virus.”
While these are not releases into the community, researchers in BSL2 labs are at a higher risk of a uuLAI than those in higher biocontainment labs or entering the community with contaminated clothing.
The CDC has issued a report[xvi] on this mix up, and the steps they have taken to avoid this particular error in the future.
The descriptions in Appendix 2 are for the eleven uuLAIs from the FOIA-requested NIH OSP incident reports. Some of the involved pathogens were not particularly virulent or contagious. If the involved pathogens were potential pandemic influenza pathogens, the outcomes could have been catastrophic. Three were due to human error, the cause of the other eight is unknown.
Appendix 1
(An enhanced metafile copy of the spreadsheet used for human error analysis)
The image in Figure A1 lists the abbreviations used for subsequent data table entries. Names of facilities have been removed as some facilities have labeled there reports as confidential. They are available, however, as they are in the public domain. The facility names are not needed for analysis of human error rates.

Figure A1. The abbreviations used for data table entries.
__________________________________________________________________________
Figure A2 on the next page shows the data collected for the first 40 incident reports.

Figure A2. First forty incidents from the FOIA NIH request. The numbers in the left-most column are for tracking the actual incident reports.
_________________________________________________________________________________
Figure A3 on the next page shows the data collected for the incident reports 41 through 80.

Figure A3. Incidents 43 through 80 from the FOIA NIH request. The numbers in the left-most column are for tracking the actual incident reports. The two red-highlighted incidents are not recorded in any data analysis for reasons stated in red-highlights
_________________________________________________________________________________
Figure A4 on the next page shows the data collected for the incident reports 80 through 123.
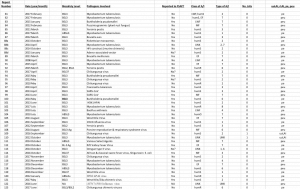
Figure A4. Incidents 81 through 123 from the FOIA NIH request. The numbers in the left-most column are for tracking the actual incident reports.
_________________________________________________________________________________
The actual spreadsheet is available from the author to anyone who wishes to analyze the data.
Appendix 2
(Detailed descriptions of the 11 uuLAIs from the FOIA NIH incident reports)
The descriptions below are for the eleven uuLAIs from the FOIA-requested NIH OSP incident reports. Many of the involved pathogens were not particularly virulent or contagious. If the involved pathogens were potential pandemic influenza pathogens, the outcomes could have been catastrophic.
Report #2. November 2004 (uuLAIs=3)
Laboratory researchers believed they were working with the [attenuated] Live Vaccine Strain (LVS) of Francisella tularensis. The LVS stock used by the researchers was contaminated with Type A F. tularensis, a wild-type, virulent form of the organism. The source of the virulent strain has not been identified to date. Three cases of Tularemia (1 confirmed, 2-probable) were identified, when three researchers had become ill in 2004 (two in May and one in September) with symptoms consistent with pneumonic tularemia. Serologic testing confirmed antibodies to F. tularensis. The experiments required BSL3, but researchers did not know they were working with an infectious strain so were working in a BSL2 lab.
The investigation into the source of the wild-type strain is ongoing.
Report #30. October 2009 (uuLAIs=1 )
A researcher acquired a laboratory Neisseria meningitides infection. The university’s ad hoc committee’s conclusion regarding the root cause of this exposure was that the researcher did not remove and dispose of his gloves after directly handling opened sample containers in the biosafety cabinet (BSC) and subsequently touched his face with the contaminated glove. Also, there is the possibility that the researcher may have been exposed to aerosol during centrifugation of samples. The researcher was subsequently placed on an IV antibiotic regimen for suspected septicemia. Personnel who may have had casual contact with the researcher were placed on a “symptom watch.”
This researcher’s inconsistent donning of appropriate personal protective equipment (PPE) while performing research was only discovered during the course of the investigation into his laboratory-acquired infection. The Institutional Biosafety Committee temporarily suspended research on biological agents in the laboratory; the Principal Investigator was allowed to continue with research not involving Neisseria meningitides. The PI indicated that the researcher was not going to conduct lab work. The researcher was instructed to work 100% time on a manuscript that is in preparation.
Report #10. April 2006 (uuLAIs=3)
Three lab workers were found to have high Coxiella Burnetti [antibody] titers. (Base line titers are performed annually.) This agent causes Q-fever. All three individuals were offered prophylaxis. None of individuals involved could recall any incident in the laboratory where exposure could have occurred and none had any clinical signs of illness.
The Biological Safety Officer made several recommendations, including following up with the Occupational Health Program, decontaminating the laboratory, and reviewing the laboratory safety procedures, including the use of personal protective equipment. The IBC was kept informed of the incident and subsequent investigation.
When they became aware of the infections, the university did not conduct a thorough investigation into their cause. Because this was not done, it was impossible to determine the precise source of exposure. The university nonetheless does not believe the titers were indicative of a laboratory exposure or infection.
[Comment: It would seem highly unusual that three laboratory workers researching Coxiella Burnetti would have become infected anywhere else.]
Report #63. August 2012 (uuLAIs=1?)
There was a potential exposure to recombinant Mycobacterium tuberculosis. The individual involved tested positive for TB in August 2012, having tested negative in a routine screening in September 2011. The individual works as a laboratory manager in the BL3 suite. While the duties and responsibilities of this position require the individual to enter and work in the BL3 lab, this person does not work directly with or handle mycobacteria. No overt exposure events were reported nor has any possible avenue for exposure in the lab been suggested; however, it cannot be ruled out that the source of exposure might be the BL3 suite.
Officials proceeded under the assumption that the lab might be contaminated and will ensure that all surfaces and equipment are carefully cleaned using an effective tuberculocidal disinfectant, per their SOPs. A retest of this individual by a different reference laboratory were all negative. It certainly appears that the test results (IGRA[xvii] only) obtained from the first lab were falsely positive.
[Comment: This incident was not included in the uuLAI data analysis.]
Report #76. September 2012 (uuLAI=1)
A lab member recently went to the Occupational Health Clinic for his semi-annual T-spot test[xviii] for Mycobacterium tuberculosis and received a positive result. There has been no known exposure or spill in the lab to cause this conversion. The lab member always wore full PPE in the BSL3 facility which included closed front gown, booties, double gloves and a Powered Air Purifying Respirator (PAPR). All procedures are conducted within a biosafety cabinet and there have no known mechanical failures related to labs HV AC or equipment. This lab member is known to be exceptionally meticulous.
Other members of the lab have been advised to consult with Occupational Health. However, several other members were tested around the same time as this individual and no others were positive. We believe this to be an isolated incident. Based on our evaluation of the conversion, we were unable to find a direct exposure or root cause of the positive test. However, as the individual’s highest risk for acquiring TB during the last 6 months was from the lab, we must assume that this a lab acquired infection. The individual is being treated by our Occupational Health Clinic.
Report # 110. January 2012 (uuLAI=1)
A potential laboratory-acquired infection (LAI) of a medical school researcher. The researcher had a positive result from a purified protein derivative (PPD) tuberculosis test taken October 1, 2012. A chest X-ray and a T-spot test were performed later that day. The chest X-ray was negative, but the T-spot test came back positive. Based on these results, a diagnosis of latent tuberculosis was made. The researcher’s previous PPD test was negative, indicating that the exposure to Mycobacterium tuberculosis occurred in the past year.
After discussing possible routes of exposure with the researcher, it was determined that the only opportunity for the researcher to have been exposed to Mycobacterium tuberculosis was when the BL3 facility experienced a power outage in December 2011. At the time of that incident, the researcher was working with Mycobacterium tuberculosis in a biosafety cabinet. However, the researcher was wearing all appropriate personal protective equipment, including an N95 respirator. No other instances of potential or overt exposure could be identified by the researcher. While it is possible that the Mycobacterium tuberculosis infection was community acquired, the University of Massachusetts Medical School is treating this as a potential LAI.
Report # 36b. March 2009 (uuLAI=1, additional tests for TB confirmed exposure)
A student who works in BSL3 containment with M tuberculosis tested positive in a routine bi-annual PPD test[xix] after repeatedly testing negative over the course of several years. The student did not have a spill, accident, PPE tear, or any other incident in the BSL3 lab that could cause obvious exposure to M tuberculosis.
The underlying cause of the PPD positive test result is therefore unknown. Prior to testing positive, the student was classified as somewhat reactive with the PPD test. It is therefore possible that repeated PPD tests over the course of several years caused the student to have an escalated response leading to what is now a positive result. Alternatively, the student may have been exposed to M tuberculosis in the outside environment. This report is being filed in the case that exposure did occur in the BLS3 lab in a manner that was not noticed.
The student was advised to seek medical consultation. The student met with a campus medical professional, given additional tests for M tuberculosis immunoreactivity, and counseled regarding options for prophylactic drug treatment. The student opted to take a course of prophylactic antibiotics as recommended. The student has not experienced any health problems.
Report # 109. December 2016 (uuLAI=1)
[Comment: This is a case of an unreported LAI, not an undetected one; thus, details about how the infection occurred were available.]
A graduate student grazed her finger with a needle while administering an antibody to mice infected with Chikungunya virus (CHIKV). She was wearing appropriate PPE. Including double gloves, but the needle broke through both pairs of gloves. She immediately washed her hands with soap and water. The student did not see any blood from the scratch so she did not report it or seek medical attention. Two days after the incident, the student developed a fever with severe body aches. Three days later, she presented with a macular rash which worsened throughout the day. That evening she reported her symptoms and the needle stick to the principal investigator (PI) and went to the hospital. She was kept in hospital overnight for observation, and the following day was seen by an infectious disease specialist who sent blood to the state laboratories for CHIKV testing. She was released from hospital that day. A few days later, the fever and rash had gone and the student did not develop arthralgia or arthritis which is often associated with Chikungunya fever. However, she did receive positive CHIKV qPCR results.
In response to this incident, the PI met with all laboratory personnel to discuss the proper reporting of personnel exposures and the processes in place for reporting incidents. The Department of Environmental Health and Safety will add additional slides about sharps safety to the annual laboratory training. The student may require additional hands on training on how to safely handle sharps. Needles should be discarded in an appropriate sharps container immediately after use to minimize the potential for a stick. The use of safety needles where the needle can be sheathed immediately after use should also be considered.
Notes
[i] Gryphon Scientific.com. Risk and Benefit Analysis of Gain of Function Research, Final Report—April 2016 (http://www.gryphonscientific.com/wp-content/uploads/2016/04/Risk-and-Benefit-Analysis-of-Gain-of-Function-Research-Final-Report.pdf)
[ii] Ibid, Table 14.1, page 507
[iii] The Department of Agriculture and the Department of Health and Human Services Report to Congress on Thefts, Losses, or Releases of Select Agents or Toxins. The reports cover the years 2003 through 2015, but only the data from 2009 through 2015 was reported in a way useful for analysis. The reports were provided to The Black Vault by the CDC under FOIA. The Black Vault is a non-government clearing house for FOIA documents. (http://www.theblackvault.com/documentarchive/department-agriculture-department-health-human-services-report-congress-thefts-losses-releases-select-agents-toxins/ ).
[iv] GAO, HIGH-CONTAINMENT LABORATORIES: Coordinated Actions Needed to Enhance the Select Agent Program’s Oversight of Hazardous Pathogens, Figure 1, United States Government Accountability Office, October 2017 (http://www.gao.gov/assets/690/687868.pdf)
[v] Select Agents and Toxins Theft, Loss or Release Information Document. Scenario 5: “A worker is manipulating a haemorrhagic fever virus suspension in a BSL-4 suit laboratory. During transport of the suspension from a biological safety cabinet to an incubator, the suspension is dropped and spills on the floor. The worker reports the incident to the supervisor, and the spill is cleaned up following established protocols. All building containment systems were operating within normal limits, and no breaches of containment suits were reported during the incident and resulting cleanup. Is this reportable?” “No,” not reportable. “The BSL-4 engineering controls in place prevented the release of the agent to the environment and exposure to the workers. However, any event that results in a breach of the positive pressure suit or secondary containment barrier should be reported to DSAT.” LK comment: Presumably “No” holds for BSL3 as well. https://www.selectagents.gov/resources/CompleteTHEFT%20LOSS%20%20RELEASE%20guidance%20document%20June82010_FINAL.pdf
[vi] NIH Guidelines for Research Involving Recombinant or Synthetic Nucleic Acid Molecules (NIH Guidelines). U.S. Department of Health and Human Services, National Institutes of Health. April 2016. Appendix G-II-C-5-a-(3). Reporting of all spills and accidents, even if relatively minor, is required as described in Appendix G-II-C-2-q. p78
[vii] There is one exception, a single incident report from 1995 where an Associate Research Scientist had set up a BSL3 experiment outside of the containment suite. The next report was in 2004.
[viii] HIGH-CONTAINMENT LABORATORIES: Improved Oversight of Dangerous Pathogens Needed to Mitigate Risk. United States Government Accountability Office, August 2016. https://www.gao.gov/assets/680/679392.pdf
[ix] ibid
[x] 2W. Liang, T. Zhao, Z. Liu, B. Guan, X. He, M. Liu, Q. Chen, G. Liu, J. Wu, R. Huang, X. Xie, and Z. Wu, “Severe Acute Respiratory Syndrome-Retrospect and Lessons of 2004 Outbreak in China,” Biomedical and Environmental Sciences, 19, (2006): 445-451.
[xi] A.C. Shurtleff, et al., The Impact of Regulations, Safety Considerations and Physical Limitations on Research Progress at Maximum Biocontainment. Viruses 2012, 4, 3932-3951; doi:10.3390/v4123932. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3528297/
[xii] Personal communication with an expert on the disbanded Soviet bioweapons program. The published source is
Никифоров В. В., ТуровскийЮ. И., Калинин П. П., АкинфееваЛ. А., Каткова Л. Р., БарминВ. С.,
Рябчикова Е. И., ПопковаН. И., Шестопалов А. М., НазаровВ. П., Ведищев С. В., Нетесов С.
В.[Nikiforov V. V., Turovskii Yu. I., Kalinin P. P., AkinfeyevaL. A., Katkova L. R., Barmin V. S.,
RyabchikovaYe. I., Popkova N. I., Shestopalov A. M., NazarovV. P., Vedishchev S. V., Netyosov S. V.]
(1994) СЛУЧАЙ ЛАБОРАТОРНОГО ЗАРАЖЕНИЯ ЛИХОРАДКОЙ МАРБУРГ. Журнал
i Immunobiologii (Moscow)] (3): 104-106 [Russian]
[xiii] Foot and Mouth Disease: Government Statement in response to investigations into the probable release of FMD virus from Pirbright. http://webarchive.nationalarchives.gov.uk/20081106100151/http://www.defra.gov.uk/animalh/diseases/fmd/investigations/pdf/govstatement_fmd2007.pdf
[xiv] http://www.bu.edu/neidl/about/construction/
[xv] HIGH-CONTAINMENT LABORATORIES. op cit. https://www.gao.gov/assets/680/679392.pdf
[xvi] Report on the Potential Exposure to Ebola Virus. Centers for Disease Control and Prevention 2/4/2015 https://stacks.cdc.gov/view/cdc/27408
[xvii] The interferon gamma release (IGRA test) assay is an important TB blood test that helps in diagnosis of tuberculosis and accessing the effectiveness of the treatment. It does not differentiate between latent infection and active disease.
[xviii] The T-SPOT. TB test is an in vitro diagnostic test for the detection of effector T cells that have been specifically activated by Mycobacterium tuberculosis antigens and is intended for use as an aid in the diagnosis of Mycobacterium tuberculosis infection (both latent and active disease).
[xix] The PPD skin test is a method used to diagnose silent (latent) tuberculosis (TB) infection. PPD stands for purified protein derivative. The PPD skin test is not a perfect screening test. A few people infected with the bacteria that cause TB may not have a reaction. Also, diseases or medicines that weaken the immune system may cause a false-negative result.
Together, we make the world safer.
The Bulletin elevates expert voices above the noise. But as an independent nonprofit organization, our operations depend on the support of readers like you. Help us continue to deliver quality journalism that holds leaders accountable. Your support of our work at any level is important. In return, we promise our coverage will be understandable, influential, vigilant, solution-oriented, and fair-minded. Together we can make a difference.
Keywords: essential biosecurity reading
Topics: Biosecurity, Disruptive Technologies















Maybe nice scholarship but very poor understanding of the biology of infectious disease. Although at first glance the data appear to support the hypothesis of community risk due to LAI, the authors ignore the fact that few of the infections listed in Appendix 1 are directly communicable. So a tularemia infected person may indeed have gotten sick due to LAI, and that is cause for concern and remediation, but is not a risk to the community because other people cannot get infected without unusual circumstances (e.g., removing a swollen lymph node from such a person and then getting cut by… Read more »