We knocked the genes out of a zebrafish—and other tales from CRISPR summer school
By Dan Drollette Jr | July 22, 2021
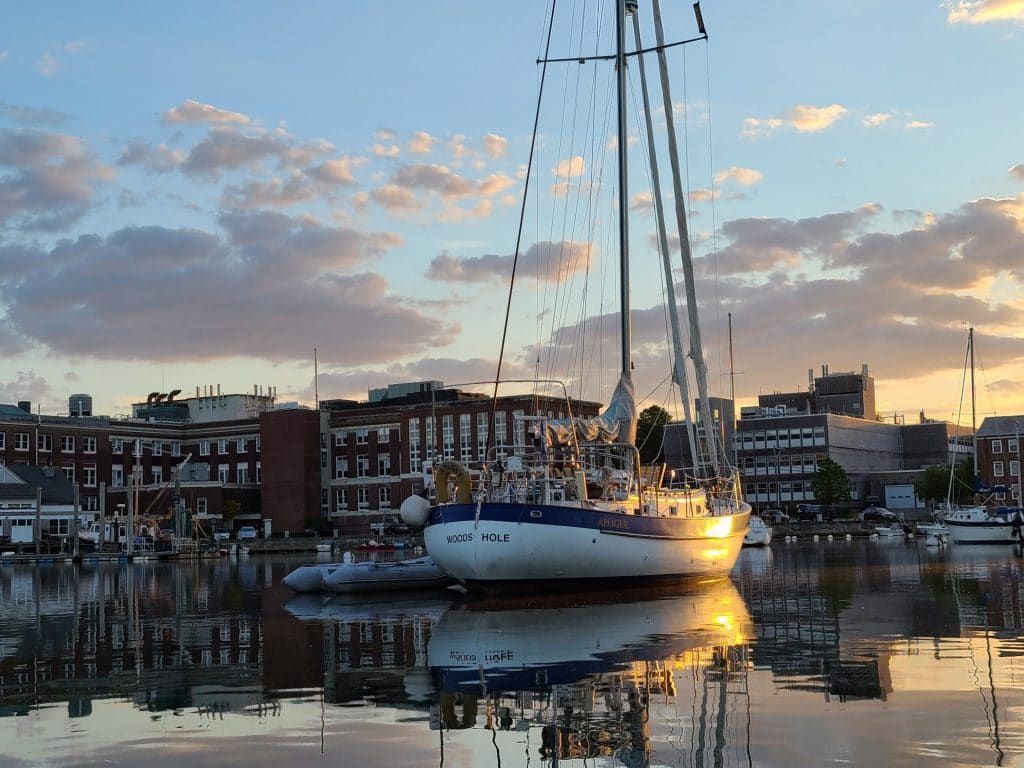 View from Eel Pond, in the center of the campus of the Marine Biological Laboratory in Woods Hole, Massachusetts. All images by Dan Drollette Jr., unless otherwise credited.
View from Eel Pond, in the center of the campus of the Marine Biological Laboratory in Woods Hole, Massachusetts. All images by Dan Drollette Jr., unless otherwise credited.
“You’re making a fish without eyeballs? What are you doing—creating some sort of Franken-fish?” That was the first response I got—which happened to come from an investigative reporter at USA Today—upon delivering the news that I was a biomedical fellow at the Logan Science Journalism Program at the Marine Biological Laboratory in Woods Hole, Massachusetts.
The second comment—also from a journalist friend, but in regard to removing the creature’s colorful spots—was: “Great. You’re transforming zebrafish into boring old guppies.”
Neither was exactly true.
Yes, I was a science journalist enrolled in a nearly two-week biomedical program last month—along with four others—at this renowned center of biological research, to get hands-on instruction in the laboratory from working biologists on how genetic editing tools like CRISPR are used.
But no, we weren’t creating Frankenstein’s monsters in fish form.
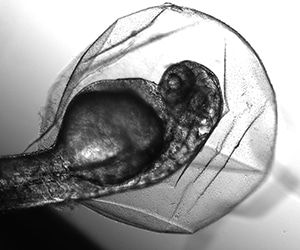
Instead, we were going to inject a tiny amount of an enzyme into the fertilized egg cells of zebrafish (Danio rerio) to see if we could stop the development of structures that are the precursors of zebrafish eyeballs—and at the same time, in a second experiment, see if we could make these fish lose their colorful markings. “CRISPR is the most powerful biological tool in human history,” Woods Hole scientist Josh Rosenthal told us on the day we started, and we journalists were going to get a better sense of what that tool could do by taking a shot at doing some bioengineering ourselves. As part of our studies, we would disable, or precisely “knock out,” some of the genes in test specimens, including those for eyeball formation and body color. If we succeeded, then these tiny, three-day-old embryonic cells, each not much more than the size of a largish grain of sand, would have no eyeball structures and no color. To confirm what we were seeing, these fish would then be analyzed on the genetic level. (For the record, none of these changes would be passed on to further generations of zebrafish. We were not working in what is called the “germ line.”)
And in a completely unrelated, third experiment in a wholly different area that had nothing at all to do with fish, we were also going to get images, using fluorescent tags and advanced optical microscopy, of the different kinds of microbial communities inside the human mouth. To do so, each of us would not brush our teeth for 24 hours and see the resulting bacteria that showed up with the aid of fluorescent imaging and a $700,000 microscope system. The DNA of each type of bacteria is unique, and we were to attach to each of them a sort of optical barcode—a short piece of synthetic DNA that recognizes a specific target microbe and reports its presence with a particular color. (See photo below.)
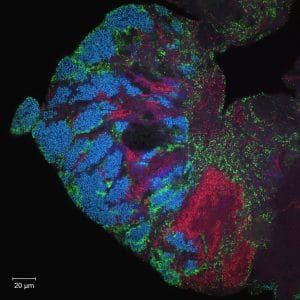
The things we do in the name of journalism.
In general, this experience was meant to give the participating journalists a more accurate and up-to-date understanding of the latest developments in bioengineering technology, how it works, and what its implications could be: Was CRISPR offering to eradicate long-term, inherited scourges? Or could its misuse be an existential threat in and of itself? Overall, was this new scientific research tool a positive, a negative … or neutral? The training would then, in theory, lead to better coverage of this complex issue in the popular press.
Why zebrafish? These minnow-like creatures are rapidly becoming one of the most desirable of lab animals for medical research. Native to the less-than-pristine waters of India’s Ganges River, these fish are hardy, easy to raise, small, and nearly transparent when young (so you can see their organs inside). A big bonus is that they mature rapidly—an important feature, because it means researchers can get results quickly. Zebrafish also share about 70 percent of their genetic code with homo sapiens, which makes it likely that the things we discover about zebrafish will have useful implications for humans.

Location, location, location. Woods Hole’s Marine Biological Laboratory has been around since 1888, making it one of the world’s oldest such institutions. “Major discoveries of fundamental biological processes—many important to human health and medical research—have come from studies of marine organisms…” says the National Academy of Sciences on its website, citing how everything from sea urchins to starfish have been used in laboratories around the world. (And yep, they call them starfish and not sea stars.) Sea creatures have features that make them great lab specimens; for example, the axons, or nerve cells, of a squid are huge and therefore easier to study than those of a human—almost the width of the lead of a pencil for a squid, or hundreds of times the width of axons in humans.
Being located on Cape Cod means that there’s an abundance of specimens nearby to collect; Woods Hole happens to be where the cold-water species of the north meet the fauna and flora of the warm waters of the Gulf Stream. And from the earliest days, it also helped that Woods Hole is situated in desirable environs, close to the ferries that service Martha’s Vineyard and other islands and within easy traveling distance of Harvard, MIT, and other Cambridge-area universities. The lab’s unique science journalism program has been around for 34 years; attendees have gone on to write for publications such as MIT’s Technology Review, the peer-reviewed journal Science, and many others.
The journalists were in a sense “embedded” with biologists and genetic engineers, closely observing what they do for a good chunk of time—and, of course, developing sources and leads to stories. Researchers tend to be so heavily involved in their specialty that they are often unaware of what makes for good fodder for an article aimed at everyday readers; they take for granted things the rest of us find extraordinary. Or as one of our instructors, Marine Biological Laboratory associate scientist Jessica Mark Welch—who led the experiment on glow-in-the-dark mouth bacteria—said about labwork: “You can forget it’s weird.”

Learning to disrobe. Security at the lab was tight; everywhere we went, we had to wear our photo IDs with magnetic swipe card. The training was thorough; six closely typed pages in the lab manual were devoted just to drawing liquid into a pipette, and we all had to take a one-hour, online lab safety course that included very specific instructions about taking off a laboratory outfit: You disrobe by removing your rubber gloves first—taking care not to touch anything with bare hands—then take off the disposable facemask, then the safety glasses, then lab coat, and then put back on a cloth face mask. There was a similar video on the details of how to dispose of dead lab specimens: sealed in a special plastic bag made for the purpose with the date, our names, and the name of the lab supervisor written on the exterior, then put in a sealed container destined for either the deep-freeze or the incinerator. My feeling was that the administration wanted to make darn sure that no species got out.

On arrival, my first impression was that the campus buildings were sort of nondescript—standard concrete and brick collegiate dorms and labs, if with the exotic feature of a dock at the end of nearly every waterside building. But nearly every building did have spectacular views of the “Eel Pond” that makes up the center of the campus; on the first day, I and the others literally gasped when we first encountered the view of the yachts in the water through the enormous picture windows that made up one entire side of our workspace in Room 306 of the Loeb Laboratory building.
Elsewhere, marine resources manager David Remsen showed us tubs that ranged in size from that of about a five-gallon bucket of paint to tanks the size of a suburban swimming pool, each filled with seawater that was carefully filtered, monitored, recycled, and heated or cooled to suit the individual requirements of the creatures under study—be they fish, sea urchins, octopus, or some other species. Seawater absolutely did not simply slop in to one side of the holding tanks and out the other, nor, Remsen insisted several times, was there any chance of an exotic invasive species sliding down the drain in the middle of the concrete floor to make an escape into the waters of the bay. (Lab escapes may have been on everyone’s mind because of the latest news about the continuing debate over the origins of the COVID-19 pandemic.)
Thankfully, we were not working with viruses and were on one of the lower danger levels when it came to biosafety, so there was no need for anything remotely like the positive pressure suits seen in photos from CDC biosafety labs. But we were doing DNA and mRNA research, knocking out genes.
A crash course. About half our time was spent with Marine Biological Laboratory senior scientist Josh Rosenthal dealing with the eyeballs and spots of zebrafish, and the other half with the Laboratory’s microbial ecologist Jessica Mark Welch studying mouth-dwelling microbes. In a typical morning or afternoon (or evening) Rosenthal, for example, would explain to us how genetic information is passed from DNA to mRNA to proteins, and how short lengths of mRNA, known as codons, can be recoded with the help of CRISPR, leading to changes in protein structure and function. In evening whiteboard lectures, we would learn about the nature of the genome and the tools and protocols used to study it—transcription, translation, primers, guides, promoters, enhancers, polymerase chain reactions, electrophoresis gels. During the day we’d actually go in the lab and prepare samples.

The emphasis was on learning and understanding the material, rather than turning us into molecular biologists. “After all, you’re here as journalists,” said Rosenthal. “So I don’t want this to be a cookbook of steps to follow. Instead, it’s more about getting down the underlying concepts.” To make the most of our time, whenever there was a lull in one experiment, we’d follow up on another. And when we were waiting for results to come back from all the experiments simultaneously, we would go out on the research vessel Gemma to collect marine samples, participate in lab tours, watch a documentary on CRISPR, attend evening lectures, and go on guided visits to places usually off-limits. We learned how sea lamprey can regenerate their spinal cords, and octopuses can camouflage themselves near-instantaneously in three-dimensions. (“I’ve got a lot of funding from the Air Force, and Army, and the Navy,” said Roger Hanlon, the head of that particular research. “I could have used camouflage like that in Vietnam.”)
Results. So, what did I learn from 10 days “embedded” with biologists at Woods Hole? A quick hit-list of impressions:
- “Marine science is gross, messy, and wet,” one of my classmates, Molly Enking, a digital editor/producer for PBS Newshour Weekend, correctly noted.
- Biology is dictated by the life-cycle of your lab animals. For example, zebrafish only breed once a day, around 8 a.m., and there’s only about a 45-minute window afterward to inject their fertilized eggs with test enzymes. Miss that opportunity, and you’ve got to wait another full day until the zebrafish reach their magical mating hour again.
- The rigid tyranny imposed by the lab animal life-cycle continues all the way down the line. Zebrafish eggs take roughly 24 hours to reach the point where they’ve developed enough for their organs to appear when you put their transparent, fertilized eggs under a microscope—and you have to do your work right then and there. (And zebrafish have not heard of weekends!)
- The outcome of one experimental step affects the next, like an endless row of dominos, until you finally get your results back from a New Jersey-based specialty processing facility in the form of electophoresis gels—thin sheets of gelatin about two by three feet in dimension, which separate out molecules by size and ultimately display your changes in the form of little blobs of genetic material clustered here and there. That way, you can confirm on the genetic level that you did indeed disable the gene for the spots on a zebrafish, in addition to actually looking at the creature’s body to see if your zebrafish is “spot-less.” You hope.
- You become involved in the results. Fellow classmate and New York City-based freelance journalist James Dinneen pointed out: “As a journalist here for a learning workshop, my career isn’t resting on this result—it’s not my life’s work, there’s no grades or grants involved, it’s purely a learning experience—but I was still hanging on what came back from the lab. I can only imagine what it’s like for a full-time researcher.”
- Using genetic engineering tools like CRISPR is easy: Look into microscope, inject little needle into egg, then press on pedal to squirt in the exact amount of reagent to knock out the tyrosinase gene responsible for the colored spots of zebrafish…
- …but not that easy. Despite the way CRISPR technology is often written up in the popular press, it’s a demanding technology, requiring patience, precision, and a capacity to deal with delayed gratification. “It also requires faith that the experiment will work,” commented classmate Alexa Kurzius of Newsela. More often than not, the experiment just doesn’t work. (My own efforts to create spot-less zebrafish failed. Though in my defense, I later learned that while I as a journalist had 10 eggs to practice on in this mock bioengineering scenario, a real-world sample was 200 eggs.) Ultimately, out of our whole group of five journalists, we got two zebrafish eggs to do what we expected—no eyeball formations in the embryos. We did better with making another set of zebrafish lose their spots; we produced a half-dozen such mosquito-larvae sized fish. But it was a valuable lesson that showed us how science offers unparalleled opportunities for disappointment. As Jessica Mark Welch noted: “Lab work is a game of constantly checking yourself; 90 percent of the time, something goes wrong.” Which is why she likes to write each step out in detail, in checklist form.

There are so many steps involved, all of which have to go exactly right, and it’s so easy for simple human error to slip in, from any number of places. As in: “Oops, was this pipette I just filled up with reagent—out of the half-dozen pipettes on the lab bench—calibrated in micro-liters or milli-liters?” And the needles are so darn tiny; you need a microscope just to see the tip of one (which you are then supposed to snap off at the very end with tweezers without breaking the rest of the ^%$#@! needle). Complicating matters, word processing programs sometimes convert the symbol “µ” for “micro” into the lowercase letter “m” for “milli”—a thousand-fold difference.
The craft of bioengineering is an aspect I had never appreciated before; the amount of effort that goes in to generate just one lab result, or “data point,” is phenomenal. This was a revelation to me, speaking as someone who had worked as the editor of a magazine put out by CERN about what is known as “high-throughput” computing. There, physicists were trying to winnow out literally billions of data points at a time.
Whereas now, I was seeing all the effort involved in getting just one data point.
Seeing all the craft and artistry that went into a simplified form of genetic engineering gave me more appreciation for the COVID-may-have-simply-escaped-from-a lab hypothesis. Stuff happens: The origin of COVID’s release could lie in something as routine and everyday as a researcher going into a bat-filled cave 600 miles away to collect samples of dung and then failing to scour herself or himself thoroughly enough upon getting back to virology headquarters.
Overall, it was a good learning experience, in a fantastic natural setting—if a bit exhausting. (Rosenthal was to tell us later that in 10 days, we had gotten about six months of graduate material on bioengineering.) On the final day of this crash course, we had to give a formal, hour-long group scientific presentation on what we learned from our experiments in the Cornelia Clapp Auditorium, the same event space where all the big-name lectures are given at the Marine Biological Laboratory, with oil paintings of scientists on the wall, and hand-made mahogany chairs on the audience floor—each containing a little brass plaque of a famous alumni, such as Rachel Carson. “To say you’ve given a talk at Clapp Auditorium is a real feather in the cap of any scientist,” said Diana Kenney, the organizer of the journalism program.
It was a bit intimidating, to say the least.

But we survived, and I returned with a better understanding of the complexities of not just the mechanics of bioengineering research but also the ethical questions involved: Would the ability to readily manipulate our genetic line lead to a dystopian brave new world, or a place free of horrific genetically inherited conditions like Huntington’s disease and sickle cell anemia? There was much to chew on.
And I learned what the acronym CRISPR means: clustered, regularly interspersed, short, palindromic, repeats.

Together, we make the world safer.
The Bulletin elevates expert voices above the noise. But as an independent nonprofit organization, our operations depend on the support of readers like you. Help us continue to deliver quality journalism that holds leaders accountable. Your support of our work at any level is important. In return, we promise our coverage will be understandable, influential, vigilant, solution-oriented, and fair-minded. Together we can make a difference.














